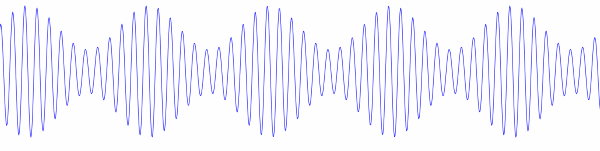
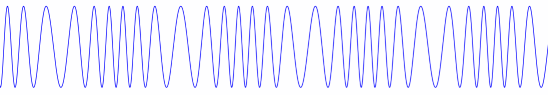
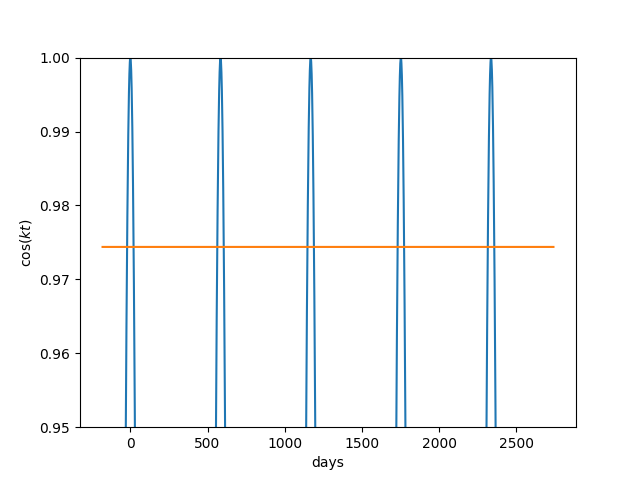
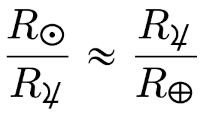Before Copernicus promoted the heliocentric model of the solar system, astronomers added epicycle on top of epicycle, creating ever more complex models of the solar system. The term epicycle is often used derisively to mean something ad hoc and unnecessarily complex.
Copernicus’ model was simpler, but it was less accurate. The increasingly complex models before Copernicus were refinements. They were not ad hoc, nor were they unnecessarily complex, if you must center your coordinate system on Earth.
It’s easy to draw the wrong conclusion from Copernicus, and from any number of other scientists who were able to greatly simplify a previous model. One could be led to believe that whenever something is too complicated, there must be a simpler approach. Sometimes there is, and sometimes there isn’t.
If there isn’t a simpler model, the time spent searching for one is wasted. If there is a simpler model, the time searching for one might still be wasted. Pursuing brute force progress might lead to a simpler model faster than pursuing a simpler model directly.
It all depends. Of course it’s wise to spend at least some time looking for a simple solution. But I think we’re fed too many stories in which the hero comes up with a simpler solution by stepping back from the problem.
Most progress comes from the kind of incremental grind that doesn’t make an inspiring story for children. And when there is a drastic simplification, that simplification usually comes after grinding on a problem, not instead of grinding on it.
3Blue1Brown touches on this in this video. The video follows two hypothetical problem solvers, Alice and Bob, who attack the same problem. Alice is the clever thinker and Bob is the calculating drudge. Alice’s solution of the original problem is certainly more elegant, and more likely to be taught in a classroom. But Bob’s approach generalizes in a way that Alice’s approach, as far as we know, does not.