What to do first after scoping out and starting a data science project?
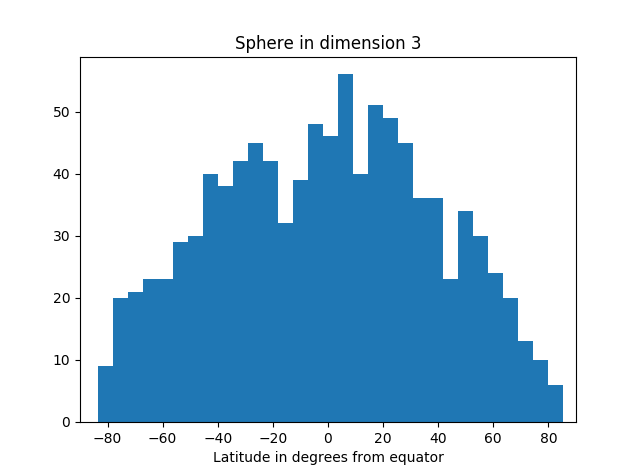
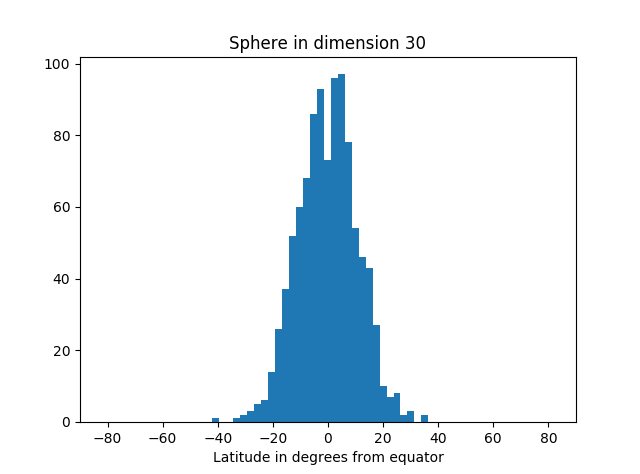
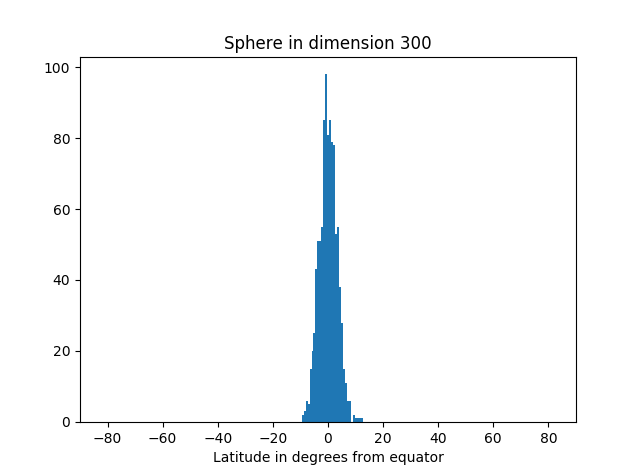
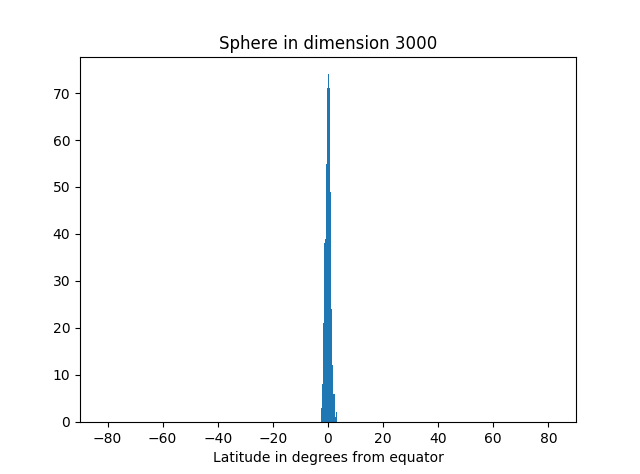
I’ve started an unsupervised learning project based on textual data. The first thing I like to do is actually look at the data. Is it noisy? What are the features—complex feature engineering needed? How heterogeneous? What generalization and overfitting challenges?
Analysis can take many forms: actually looking at the numbers, using visualization tools, Excel spreadsheet, Jupyter notebooks with Matplotlib, computing various statistics on the whole dataset or portions of it.
Some may believe this is not important. Just throw a barrage of classification or regression methods at the data, treat the data as a black box. Of course testing on a suite of ML methods is not a bad thing. But I can’t imagine not using every avenue available, including looking at the data. I’m certainly not alone in this view (see for example here, here and here).
I spent a few hours developing a simple custom data viewer for my problem that colored different parts of the textual data to give insight as to what was going on. I used ChatGPT to develop parts of this tool; some of it was incorrect and needed fixing, but having at least a draft of the code definitely saved time. Seeing the actual data in person was insightful and generated ideas for solving the problem.
While inspecting the data can help identify issues, it also risks biasing the modeling process by imposing assumptions that a flexible model might otherwise uncover on its own. One must also beware of data leakage. That being said—in general I think understanding as much as you can about the data is not a bad thing.