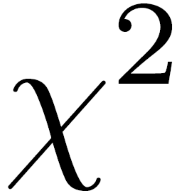
Yesterday Brian Hayes wrote a post about the distribution of primes. He showed how you could take the remainder when primes are divided by 7 and produce something that looks like rolls of six-sided dice. Here we apply the chi-square goodness of fit test to show that the rolls are too evenly distributed to mimic randomness. This post does not assume you’ve seen the chi-square test before, so it serves as an introduction to this goodness of fit test.
In Brian Hayes’ post, he looks at the remainder when consecutive primes are divided by 7, starting with 11. Why 11? Because it’s the smallest prime bigger than 7. Since no prime is divisible by any other prime, all the primes after 7 will have a remainder of between 1 and 6 inclusive when divided by 7. So the results are analogous to rolling six-sided dice.
The following Python code looks at prime remainders and (pseudo)random rolls of dice and computes the chi-square statistic for both.
First, we import some functions we’ll need.
from sympy import prime
from random import random
from math import ceil
The function prime takes an argument n and returns the nth prime. The function random produces a pseudorandom number between 0 and 1. The ceiling function ceil rounds its argument up to an integer. We’ll use it to convert the output of random into dice rolls.
In this example we’ll use six-sided dice, but you could change num_sides to simulate other kinds of dice. With six-sided dice, we divide by 7, and we start our primes with the fifth prime, 11.
num_sides = 6
modulus = num_sides + 1
# Find the index of the smallest prime bigger than num_sides
index = 1
while prime(index) <= modulus:
index += 1
We’re going to take a million samples and count how many times we see 1, 2, …, 6. We’ll keep track of our results in an array of length 7, wasting a little bit of space since the 0th slot will always be 0. (Because the remainder when dividing a prime by a smaller number is always positive.)
# Number of samples
N = 1000000
observed_primes = [0]*modulus
observed_random = [0]*modulus
Next we “roll” our dice two ways, using prime remainders and using a pseudorandom number generator.
for i in range(index, N+index):
m = prime(i) % modulus
observed_primes[m] += 1
m = int(ceil(random()*num_sides))
observed_random[m] += 1
The chi-square goodness of fit test depends on the observed number of events in each cell and the expected number. We expect 1/6th of the rolls to land in cell 1, 2, …, 6 for both the primes and the random numbers. But in a general application of the chi-square test, you could have a different expected number of observations in each cell.
expected = [N/num_sides for i in range(1, modulus)]
The chi-square test statistic sums (O – E)2/E over all cells, where O stands for “observed” and E stands for “expected.”
def chisq_stat(O, E):
return sum( (o - e)**2/e for (o, e) in zip(O, E) )
Finally, we compute the chi-square statistic for both methods.
ch = chisq_stat(observed_primes[1:], expected[1:])
print(ch)
ch = chisq_stat(observed_random[1:], expected[1:])
print(ch)
Note that we chop off the first element of the observed and expected lists to get rid of the 0th element that we didn’t use.
When I ran this I got 0.01865 for the prime method and 5.0243 for the random method. Your results for the prime method should be the same, though you might have a different result for the random method.
Now, how do we interpret these results? Since we have six possible outcomes, our test statistics has a chi-square distribution with five degrees of freedom. It’s one less than the number of possibilities because the total counts have to sum to N; if you know how many times 1, 2, 3, 4, and 5 came up, you can calculate how many times 6 came up.
A chi-square distribution with ν degrees of freedom has expected value ν. In our case, we expect a value around 5, and so the chi-square value of 5.0243 is unremarkable. But the value of 0.01864 is remarkably small. A large chi-square statistics would indicate a poor fit, the observed numbers being suspiciously far from their expected values. But a small chi-square value suggests the fit is suspiciously good, closer to the expected values than we’d expect of a random process.
We can be precise about how common or unusual a chi-square statistic is by computing the probability that a sample from the chi square distribution would be larger or smaller. The cdf gives the probability of seeing a value this small or smaller, i.e. a fit this good or better. The sf gives the probability of seeing a value this larger or larger, i.e. a fit this bad or worse. (The scipy library uses sf for “survival function,” another name for the ccdf, complementary cumulative distribution function).
from scipy.stats import chi2
print(chi2.cdf(ch, num_sides-1), chi2.sf(ch, num_sides-1))
This says that for the random rolls, there’s about a 41% chance of seeing a better fit and a 59% chance of seeing a worse fit. Unremarkable.
But it says there’s only a 2.5 in a million chance of seeing a better fit than we get with prime numbers. The fit is suspiciously good. In a sense this is not surprising: prime numbers are not random! And yet in another sense it is surprising since there’s a heuristic that says primes act like random numbers unless there’s a good reason why in some context they don’t. This departure from randomness is the subject of research published just this year.
If you look at dice with 4 or 12 sides, you get a suspiciously good fit, but not as suspicious as with 6 sides. But with 8 or 20-sided dice you get a very bad fit, so bad that its probability underflows to 0. This is because the corresponding moduli, 9 and 21, are composite, which means some of the cells in our chi-square test will have no observations. (Suppose m has a proper factor a. Then if a prime p were congruent to a mod m, p would be have to be divisible by a.)
Update: See the next post for a systematic look at different moduli.
You don’t have to use “dice” that correspond to regular solids. You could consider 10-sided “dice,” for example. For such numbers it may be easier to think of spinners than dice, a spinner with 10 equal arc segments it could fall into.
Related post: Probability that a number is prime